Our brain consists of a labyrinth of neurons that communicate with each other via electrical signals, i.e. voltage pulses. Neuroscientists aim to understand how such neuronal circuits modulate brain function. For this purpose, it is extremely helpful to visualize voltage changes at the membrane of nerve cells. A promising possibility to make voltages literally visible is offered by certain sensors, the so-called genetically encoded voltage indicators (GEVIs). These GEVI sensors are proteins that sit in the cell membrane where they can sense the membrane potential and relate a change in voltage to a form of output, often fluorescence. GEVIs are powerful tools, that can be used to image large neuronal networks or even individual cells. The search for new, advanced GEVI sensors is an active field of research.
In their recent study, a team involving the UniSysCat groups of Peter Hildebrandt, Han Sun, Tillmann Utesch, and Peter Hegemann gives new insights into the function of a certain class of GEVIs: They studied variants (e.g. QuasArs) of the microbial protein archaerhodopsin-3 (Arch3). In contrast to other rhodopsins, these variants are fluorescent upon changes in transmembrane voltage and have already been used for imaging membrane voltage changes in cell cultures and small animals.
Arch3-based voltage indicators could be particularly valuable in neuroscience, where they are combined with optogenetic actuators and provide a non-invasive monitoring of the membrane voltage dynamics. However, so far, these sensors fluoresce only very weakly. The highest reported fluorescence quantum yield (QY) of Arch3-based voltage sensors is still only ~1.2% whereas the highest reported rhodopsin fluorescence QY reaches 20%. Hence, it is highly desirable to further improve the performance of Arch3-based sensors for applications in neuroscience.
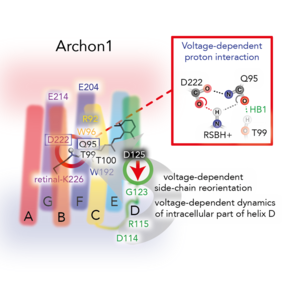
In their current paper, the authors apply a combination of spectroscopy studies, imaging techniques, and atomistic molecular dynamics (MD) simulations to investigate various Arch3 variants in order to understand the origin of the fluorescence and how it is controlled by the voltage. Their experiments and simulations provide substantial new insights into the fluorescence of microbial rhodopsins: The authors obtained a better understanding of the impact of the membrane voltage on the protein dynamics and the fluorescence by identifying key structural sites of the protein that control the fluorescence intensity. These insights offer now a basis to rationally develop more advanced voltage indicators with enhanced fluorescence.
This study has been published as an Open Access Paper in Nature Communications: Silapetere, A., Hwang, S., Hontani, Y. et al. QuasAr Odyssey: the origin of fluorescence and its voltage sensitivity in microbial rhodopsins. Nat Commun 13, 5501 (2022). https://doi.org/10.1038/s41467-022-33084-4